MILCDock: Machine Learning Enhanced Consensus Docking for Virtual Screening in Drug Discovery | Journal of Chemical Information and Modeling
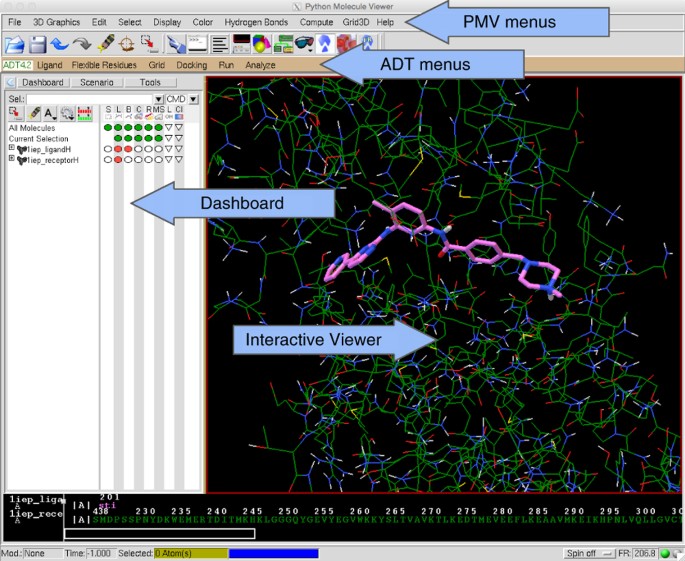
Computational protein–ligand docking and virtual drug screening with the AutoDock suite | Nature Protocols
![PDF] An extensive survey of molecular docking tools and their applications using text mining and deep curation strategies. | Semantic Scholar PDF] An extensive survey of molecular docking tools and their applications using text mining and deep curation strategies. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/875c8187625cf4e582c0c02e7f128e055e7c5c25/53-Table2-1.png)
PDF] An extensive survey of molecular docking tools and their applications using text mining and deep curation strategies. | Semantic Scholar

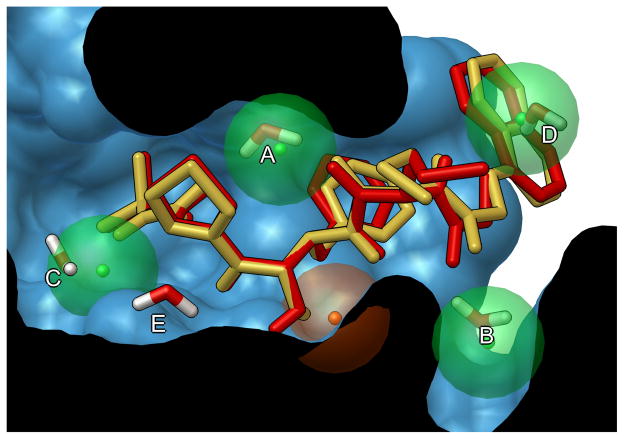
Virtual screening as a tool to discover new β-haematin inhibitors with activity against malaria parasites | Scientific Reports

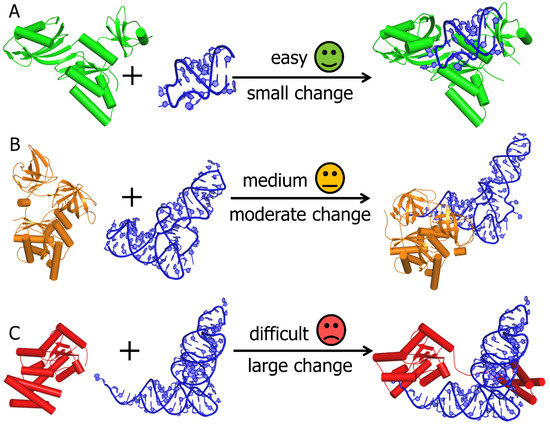
Protein-based Virtual Screening Tools applied for RNA-Ligand Docking identify new Binders of the preQ1-Riboswitch | bioRxiv

AMDock: a versatile graphical tool for assisting molecular docking with Autodock Vina and Autodock4 | Biology Direct | Full Text